One can calculate robust standard errors in R in various ways. However, one can easily reach its limit when calculating robust standard errors in R, especially when you are new in R. It always bordered me that you can calculate robust standard errors so easily in STATA, but you needed ten lines of code to compute robust standard errors in R. I decided to solve the problem myself and modified the summary() function in R so that it replicates the simple way of STATA. I added the parameter robust to the summary() function that calculates robust standard errors if one sets the parameter to true. With the new summary() function you can get robust standard errors in your usual summary() output. All you need to do is to set the robust parameter to true:
summary(lm.object, robust=T)
Furthermore, I uploaded the function to a github.com repository. This makes it easy to load the function into your R session. The following lines of code import the function into your R session. You can also download the function directly from this post yourself.
# load necessary packages for importing the function
library(RCurl)
# import the function from repository
url_robust <- "https://raw.githubusercontent.com/IsidoreBeautrelet/economictheoryblog/master/robust_summary.R"
eval(parse(text = getURL(url_robust, ssl.verifypeer = FALSE)),
envir=.GlobalEnv)
I prepared a working example that carries out an OLS estimate in R, loads the function to compute robust standard errors and shows to apply it. You find the code below.
# start with an empty workspace
rm(list=ls())
# load necessary packages for importing the function
library(RCurl)
# load necessary packages for the example
library(gdata)
library(zoo)
# import the function
url_robust <- "https://raw.githubusercontent.com/IsidoreBeautrelet/economictheoryblog/master/robust_summary.R"
eval(parse(text = getURL(url_robust, ssl.verifypeer = FALSE)),
envir=.GlobalEnv)
# download data set for example
url_data <- "https://economictheoryblog.com/wp-content/uploads/2016/08/data.xlsx"
data <- read.xls(gsub("s:",":",url_data))
# estimate simple linear model
reg <- lm(weight ~ lag_calories+lag_cycling+
I(lag_calories*lag_cycling),
data=data)
# use new summary function
summary(reg)
summary(reg,robust = T)
# strangely enough we get a case in which
# robust standard errors a smaller than
# conventional standard errors
One can also easily include the obtained robust standard errors in stargazer and create perfectly formatted tex or html tables. This post describes how one can achieve it.

The lack of the “robust” option was among my biggest disappointments in moving our courses (and students) from STATA to R. We will all be eternally grateful to you for rectifying this problem.
Thank you for your kind words of appreciation. I’m glad I was able to help.
Have you come across a heteroscedasticity-robust F-test for multiple linear restrictions in a model?
I can’t say I have.
What I know is that, once you start using heteroscedasticity consistent standard errors you should not use the sums of squares to calculate the F-statistic. The reason for this is that the meaning of those sums is no longer relevant, although the sums of squares themselves do not change. To my understanding one can still use the sums of squares to calculate the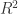 statistic that maintains its goodness-of-fit interpretation. However, you cannot use the sums of squares to obtain F-Statistics because those formulas do no longer apply. Instead of using an F-Statistic that is based on the sum of squared what one does is to use a Wald test that is based on the robustly estimated variance matrix.
statistic that maintains its goodness-of-fit interpretation. However, you cannot use the sums of squares to obtain F-Statistics because those formulas do no longer apply. Instead of using an F-Statistic that is based on the sum of squared what one does is to use a Wald test that is based on the robustly estimated variance matrix.
So, if you use my function to obtain robust standard errors it actually returns you an F-Statistic that is based on a Wald test instead of sum of squares.
I suppose that if you want to test multiple linear restrictions you should use heteroscedasticity-robust Wald statistics. I don’t know that if there is actually an R implementation of the heteroscedasticity-robust Wald.
For now I am working on an implementation of clustered standard errors, but once I am done with it I might look into it myself.
HTH
I assumed that, if you went to all the hard work to calculate the robust standard errors, the F-statistic you produced would use them and took it on faith that I had the robust F. Stock and Watson report a value for the heteroscedasticity-robust F stat with q linear restrictions but only give instructions to students for calculating the F stat under the assumption of homoscedasticy, via the SSR/R-squared (although they do describe the process for coming up with the robust F in an appendix).
Having the robust option in R is a great leap forward for my teaching. The rest can wait.
Thanks again,
Matt
Awesome! Extremely useful! Thank you..
Thanks, I am glad to hear that!
Thanks for this. I was playing with R a couple years back thinking I’d make the switch and was baffled by how difficult it was to do this simple procedure. Does this only work for lm models? I tried it with a logit and it didn’t change the standard errors.
Hi. Unfortunately, the function only covers lm models so far. However, I will extent the function to logit and plm once I can free up some time. The “sandwich” package, created and maintained by Achim Zeileis, provides some useful functionalities with respect to robust standard errors. Cheers.
Hi all, interesting function. Previously, I have been using the sandwich package to report robust S.E.s. When I installed this extension and used the summary(, robust=T) option slightly different S.E.s were reported from the ones I observed in STATA. I trimmed some of my results and posted them below. Do you know why the robust standard errors on Family_Inc don’t match ?
> coeftest(mod1, vcov = vcovHC(mod1, “HC1”)) #Robust SE (Match those reported by STATA)
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.3460131 0.0974894 24.064 < 2.2e-16 ***
Famliy_Inc 0.5551564 0.0086837 63.931 summary(mod1, robust = T) #Different S.E.s reported by robust=T
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.346013 0.088341 26.56 <2e-16 ***
Family_Inc 0.555156 0.007878 70.47 <2e-16 ***
Hi! Thank you for your interest in my function. I am surprised that the standard errors do not match. On my blog I provide a reproducible example of a linear regression with robust standard errors both in R and STATA. Both programs deliver the same robust standard errors. See the following two links if you want to check it yourself:
Furthermore, I also check coeftest(reg, vcov = vcovHC(reg, “HC1”)) for my example and the sandwich version of computing robust standard errors calculates the same values.
Unfortunately, I cannot tell you more right now. Could you provide a reproducible example? I am very keen to know what drives the differences in your case. Especially if the are a result of my function.
Cheers!
I am seeing slight differences as well. Depending on the scale of your t-values this might be a issue when recreating studies. Take this example, recreating a study by Miguel et al. (2004):
library(readstata13)
library(dplyr)
library(countrycode)
# get the data mss_repdata.dta from http://emiguel.econ.berkeley.edu/research/economic-shocks-and-civil-conflict-an-instrumental-variables-approach
df <- read.dta13(file = "mss_repdata.dta")
df$iso2c |t| [95% Conf. Interval]
# ————-+—————————————————————-
# GPCP_g | .0554296 .0163015 3.40 0.002 .0224831 .0883761
# GPCP_g_l | .0340581 .0132131 2.58 0.014 .0073535 .0607628
# _cons | -.0061467 .0024601 -2.50 0.017 -.0111188 -.0011747
# Same regression in R:
# Country specific time trends
df % group_by(ccode) %>% mutate(tt = year-1978)
for (cc in unique(df$iso2c)) {
tmp <- df[df$iso2c == cc,]$tt
df[, paste0("tt.", cc)] <- ifelse(df$iso2c == cc, tmp, 0)
}
## Country fixed effects
for (cc in unique(df$iso2c)) {
df[, paste0("fe.", cc)] <- ifelse(df$iso2c == cc, 1, 0)
}
ols |t|)
# (Intercept) -0.00615 0.00262 -2.35 0.0191 *
# GPCP_g 0.05543 0.01418 3.91 0.0001 ***
# GPCP_g_l 0.03406 0.01190 2.86 0.0043 **
# —
# The same applies for:
coeftest(ols, vcov = function(x) sandwich::vcovHC(x, type = “HC1”, cluster = “group”))
Well, code in comments is not ideal I guess. Take this git link instead: https://github.com/martinschmelzer/Miguel/blob/master/miguel_robust.R
Hey Martin! Thank you for you remark and the reproducible example. The function to compute robust standard errors in R works perfectly fine. The reason why the standard errors do not match in your example is that you mixed up some things. However, first things first, I downloaded the data you mentioned and estimated your model in both STATA 14 and R and both yield the same results. That is, if you estimate “summary.lm(lm(gdp_g ~ GPCP_g + GPCP_g_l), robust = T)” in R it leads to the same results as if you estimate “reg gdp_g GPCP_g GPCP_g_l, robust” in STATA 14. That problem is that in your example you do not estimate “reg gdp_g GPCP_g GPCP_g_l, robust” in STATA, but you rather estimate “reg gdp_g GPCP_g GPCP_g_l, cluster(country_code)”. That of course does not lead to the same results. The reason being that the first command estimates robust standard errors and the second command estimates clustered robust standard errors. Two very different things.
If you want to estimate OLS with clustered robust standard errors in R you need to specify the cluster. In your case you can simply run “summary.lm(lm(gdp_g ~ GPCP_g + GPCP_g_l), cluster = c(“country_code”))” and you obtain the same results as in your example.
Check out the instructions for clustered standard errors in R on the following post: https://economictheoryblog.com/2016/12/13/clustered-standard-errors-in-r/
Thanks again for you comment. I hope I was able to help you.
Cheers
Will I need to import this function every time start a session or will this permanently change the summary() function?
Unfortunately, you need to import the function every session. Best, ad